Assessing soil health: Measuring the soil microbiome

The following article is the fourth in a six-part series on assessing soil health. It focuses on measuring the soil microbiome. It is part of a larger Soil Science Society of America webinar series produced in partnership with The Soil Health Institute and sponsored by The Walton Family Foundation.
Soil microbial community members are responsible for maintaining a multitude of functions in soil. Microbial community members enhance soil health through transformations of organic residues to plant-available nutrients, breakdown harmful inorganic chemicals, stabilization of soil aggregates, and regulation of crop diseases.
Welcome to the fourth installment of the Assessing Soil Health series, which is all about measurements of the soil microbiome. This article will discuss insights from general soil science literature and from the recent Soil Health Institute project titled, “The North American Project to Evaluate Soil Health Measurements (NAPESHM).” Evaluations will include measures of microbial respiration, potential enzyme activity, phospholipid fatty acids (PLFAs), targeted amplicon sequencing, and metagenomic sequencing.
Soil Sampling for Microbiome Measurements
Soil microbial communities are sensitive to climate and inherent soil properties as well as agricultural management. Hence, sampling consistently in relation to management and temporal variables is important when applying measurements of the microbiome to understand how changes in management affect soil microbial communities. Variables that affect consistent sampling timing include soil temperature and moisture. Like other soil health sampling, microbiome soil samples should be taken from 0–15 cm. However, unlike other soil sampling, samples must remain cool when transported for laboratory analyses. It is recommended to keep samples for PLFA and genomic microbiome measurements cool, but not frozen, to reduce potential microbial activity capable of changing microbial community structure. Additionally, when measurements require DNA extraction, it is important to sterilize sampling equipment with ethanol in between sampling treatments.
Methods Currently Available by Commercial Laboratories
Microbial Respiration
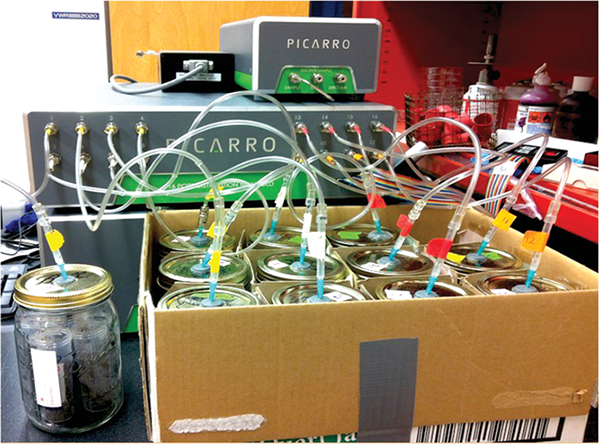

Soil microbial community members respire carbon dioxide as a byproduct of metabolism, like how most animals exhale carbon dioxide in each breath. Standardized measures of respiration quantify flushes of carbon dioxide released from microbes in rewetted soil during a short-term aerobic incubation (Figure 1). Measurements of respiration generally increase as a result of implementing soil health management systems that are building soil organic matter, which serves as a microbial food source, fueling their activity. Results from NAPESHM generally indicate increased respiration in systems with residue retention, organic amendments, cover crops, and reduced tillage. However, if a sample is taken immediately following a tillage event, there is a spike in respiration. This spike is temporary and does not reflect respiration differences between management activities throughout the majority of the year. We recommend consistent sampling timing before spring management activities whenever possible.
Phospholipid Fatty Acids
Phospholipid fatty acids are the primary component of cellular membranes, which are comprised of phospholipid heads and fatty acid tails. Measurements of PLFA are interpreted as a measure of the living microbial community in a soil because membrane components quickly degrade following the death of a cell.
Individual PLFAs are distinguished by the number of carbon atoms contained in the fatty acid tail as well as bond saturation and conformation; however, there is no standard method for accounting in PLFA analyses. For example, the NAPESHM project scientists calculated microbial biomass as the sum of PLFAs with fatty acid tail lengths from 14 to 20 carbon atoms. Several academic research groups have used similar sets of biomarkers, but private laboratories can use different combinations of PLFAs to calculate biomass. It was originally hypothesized that groups within the soil microbial community (e.g., arbuscular mycorrhizal fungi, gram positive or negative bacteria, and actinomycetes) could be distinguished by their PLFA fingerprint. However, research has shown PLFA fingerprints of individual community members may change based on environmental conditions (Frostegård et al., 2011). Our shifting understanding in interpreting PLFA fingerprints limits confidence in interpretation of PLFA data to quantify community members’ presence and prevalence in soil.
Potential Enzyme Activity
While microbial respiration provides a generalized measure of microbial activity in soil, potential enzyme activity measures a soil’s capacity to degrade specific organic molecules. The USDA-NRCS recommends measuring the potential activity of four enzymes: Beta-glucosidase, N-aceytl beta-glucosiminidase, phosphomonoesterase, and arylsulfatase. The four enzymes use similar mechanisms to hydrolyze specific bonds to break apart organic compounds. Beta-glucosidase hydrolyzes beta-glucosidic bonds in cellulose, releasing glucose molecules. N-aceytl beta-glucosiminidase catalyzes the terminal reaction in chitin degradation, the basic structural component of fungal cell walls and insect exoskelatons. Phosphomonoesterase catalyzes the release of phosphates, making it available for crop uptake. Arylsulfatase catalyzes the release of sulfate. Potential enzyme activity is measured by saturating a sample with a specific substrate and quantifying the number of molecules released. Interpretation of potential enzyme activity in relation to soil health is difficult because multiple mechanisms can drive measurement results up or down. The lack of broad interpretability limits its use as a soil health indicator at scale.
Potential DNA-Based Measurements Hold Potential for Soil Health Interpretation
The two methods highlighted below are currently being assessed by the scientific community to understand their potential applicability as soil health indicators. Both methods are based on microbial DNA extracted from soil samples. Interpretation of DNA provides information on the structure of microbial communities and their capacity to perform specific functions. However, these measurements must first be linked to functional outcomes (e.g., carbon storage and efficient nutrient transformation) prior to application as indicators.
Targeted Amplicon Sequencing
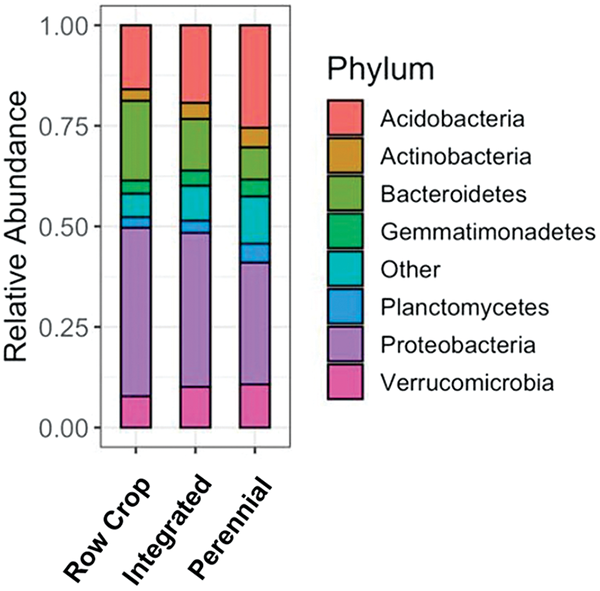
Targeted amplicon sequencing uses DNA sequences of bacterial, archaeal, and fungal origin to identify individual microbial community members. The relative number and types of bacteria and archaea in the community are measured with 16S rRNA amplicon sequencing while fungal community structure is measured with ITS and 18S rRNA. Targeted DNA sequences are extracted from soil and matched against databases to identify the presence of individual organisms. Currently, the number of sequences extracted from a soil sample is not standardized in laboratory procedures; therefore, the measurement should be interpreted in terms of relative abundances. Shifts in the relative abundances of groups of organisms provide an understanding of how management practices select for microbial community members with different life strategies. While shifts due to changes in management practices may seem small, a 1% change in microbial relative abundance is equivalent to altering the presence of hundreds of thousands of organisms. Figure 2 shows how introducing perennials into a row-cropping system shifts microbial community structure towards native perennial systems.
Metagenomic Sequencing
Metagenomic sequencing provides information on all genetic material contained in the soil. Soil DNA is aligned with gene libraries to identify specific genes that give the soil organism the potential to perform specific tasks. While potential enzyme activity is limited to interpreting a single reaction necessary for transformation of nutrients into plant-available forms, metagenomic sequencing holds this potential to understand entire transformation pathways. Like targeted amplicon sequencing, the sequences should be interpreted as relative proportions of genes in samples. To date, few peer-reviewed publications have assessed how agricultural management affects soils’ genetic capacity to function.
Summary
Soil health measurements indicate microbiome size, potential activity, and structure. While many measurements are responsive to agricultural management, the majority are difficult to link to functional outcomes, making meaningful interpretation difficult. Microbial respiration provides a general measurement of activity; however, sampling over multiple years should remain consistent with management practices, soil moisture, and soil temperature. Genomic techniques hold great potential as useful indicators of soil health, but first must be linked to functional outcomes.

References
Frostegård, Å., Tunlid, A., & Bååth, E. (2011). Use and misuse of PLFA measurements in soils. Soil Biology and Biochemistry, 43, 1621–1625. https://doi.org/10.1016/j.soilbio.2010.11.021
Text © . The authors. CC BY-NC-ND 4.0. Except where otherwise noted, images are subject to copyright. Any reuse without express permission from the copyright owner is prohibited.